Comparative Analysis Association and Prediction of Various Phenotypic Traits of Oryza Sativa
Main Article Content
Abstract
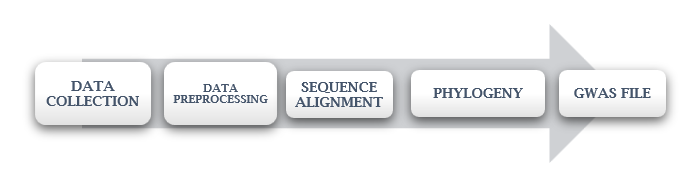
Understanding the genotype-phenotype relationship and accurately predicting breeding values are crucial aspects of crop improvement programs. This paper investigates the genetic basis ,association of phenotypic trait height and yield and predicts the phenotypic traits of Oryza Sativa (rice) through a comprehensive approach encompassing genome-wide association studies (GWAS), phylogenetic analysis, machine learning algorithms, and the development of a graphical user interface (GUI) application. Genotypic and phenotypic data were collected from the RiceVarMap database. The genotypic information consisted of gene variation IDs, while the phenotype data included plant height. Data preprocessing involved the creation of a sequence. fasta file and multiple sequence alignment using the ClustalW tool. A phylogenetic tree was then constructed to analyse the subpopulations of Oryza Sativa. Clustering techniques were applied to further explore the genetic relationships among the samples. A GWAS file was generated to identify associations between genotype and phenotype. Subsequently, machine learning algorithms were employed for the classification and prediction of genomic estimated breeding values (GEBV) for height and yield traits. Random Forest emerged as the most accurate algorithm with 85% accuracy. To facilitate user interaction and data exploration, a GUI application was developed using Flask, allowing users to access the phylogenetic tree, height, and yield information, GWAS results, and make predictions. We explored there is a strong positive association between phenotypic trait height and yield.